  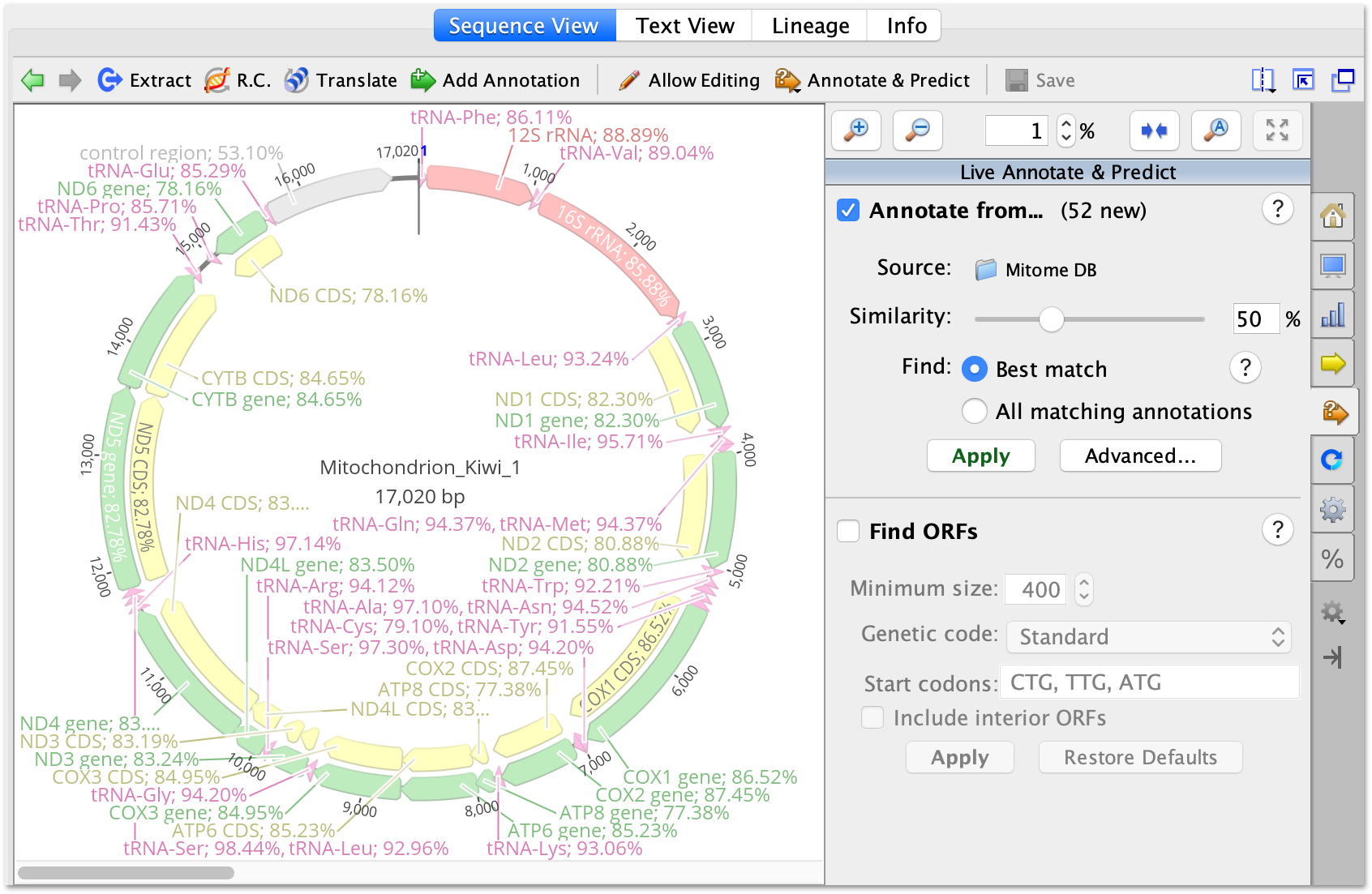
If your Geneious file contains metadata that has been added via the Sequence view Info tab then you can check the option to Include Meta-data in Comments section of GenBank File to have the metadata exported in the COMMENT section of the GenBank file. On Window Vista, Windows 7, 8 and 10 you need to run Geneious as an administrator to change this setting (close Geneious, then right-click on the Geneious icon and choose. You may need to restart for the change to take effect. To ensure maximum compatibility, uncheck the option to Allow names longer than 16 characters, and check the option to Replace spaces in names with underscores. In Geneious, go into ToolsPreferences, and under the General tab increase the Max memory available to Geneious. All annotations that do not use standard Feature names will be converted to Feature Type misc_feature. In the Export settings window check the option to Convert annotations to strict GenBank format. To export your file with Features in strict GenBank format, select the file/s to be exported, and go menu File - Export - Selected documents. However, software external to Geneious may not be able to handle non-standard feature names or non-standard locus names. Geneious can handle sequences with Annotations (Features) with non-standard Feature names, and is able to correctly import GenBank files that contain non-standard Feature names. Strict GenBank format requires that Locus name has a maximum length of 16 characters with no internal spaces, and that only standard Features (as listed in Table 2 above) are included. Geneious will correctly handle these deprecated Feature types if present.Įxporting from Geneious in Strict GenBank format † As of, regulatory replaces enhancer, promoter, CAAT_signal, TATA_signal, -35_signal, -10_signal, RBS, GC_signal, polyA_signal, attenuator, terminator, misc_signal. The functionality in Geneious Basic can be compared with a number of other desktop software packages, such as VectorNTI (Lu and Moriyama, 2004), CLC Bio. 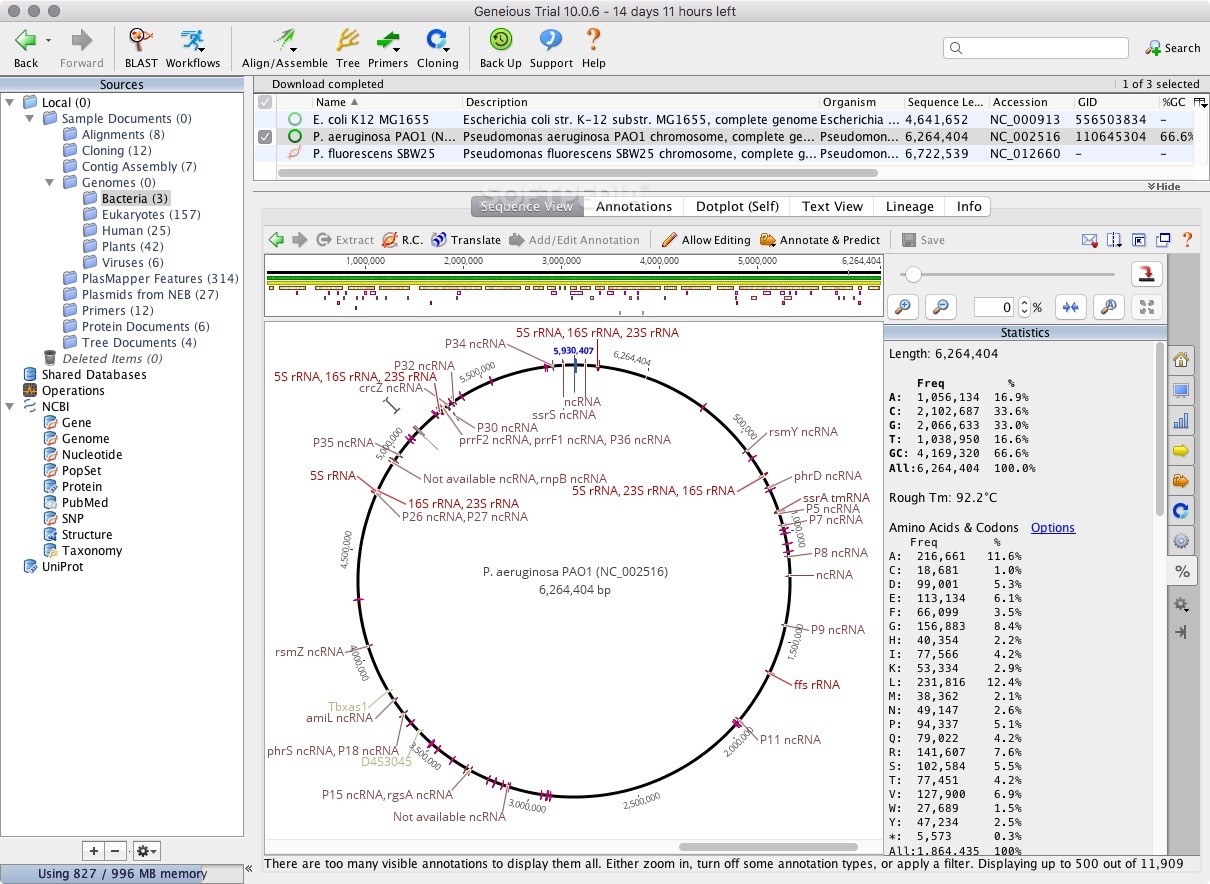
Table 2: Standard names for Features used in GenBank files. Table 2 lists standard Features names prescribed for use in GenBank Files, see for a complete summary which also describes associated qualifiers for each Feature type. Annotations are the equivalent of GenBank Features.Īnnotations will usually have associated Properties information (in GenBank referred to as Qualifiers) and Interval coordinates (In GenBank referred to as location descriptors). In Geneious Prime functional elements on DNA and protein sequence are recorded and depicted as Annotations. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
